FEATURES
GPS-Prot is the quickest and easiest way to begin exploring the questions: “What does my protein of interest bind to?” & “What are the publications and data supporting this interaction?”
- COMPREHENSIVE: more than 300,000 unique Scored Human PPIs from major curated databases (Biogrid, MINT, IntACT, DIP, HPRD, MIPS and more, as implemented in HIPPIE)
- EASY TO GET STARTED: No software to download or install. Generate a protein interaction (PPI) network for any protein simply by entering its gene ID. Requires Javascript.
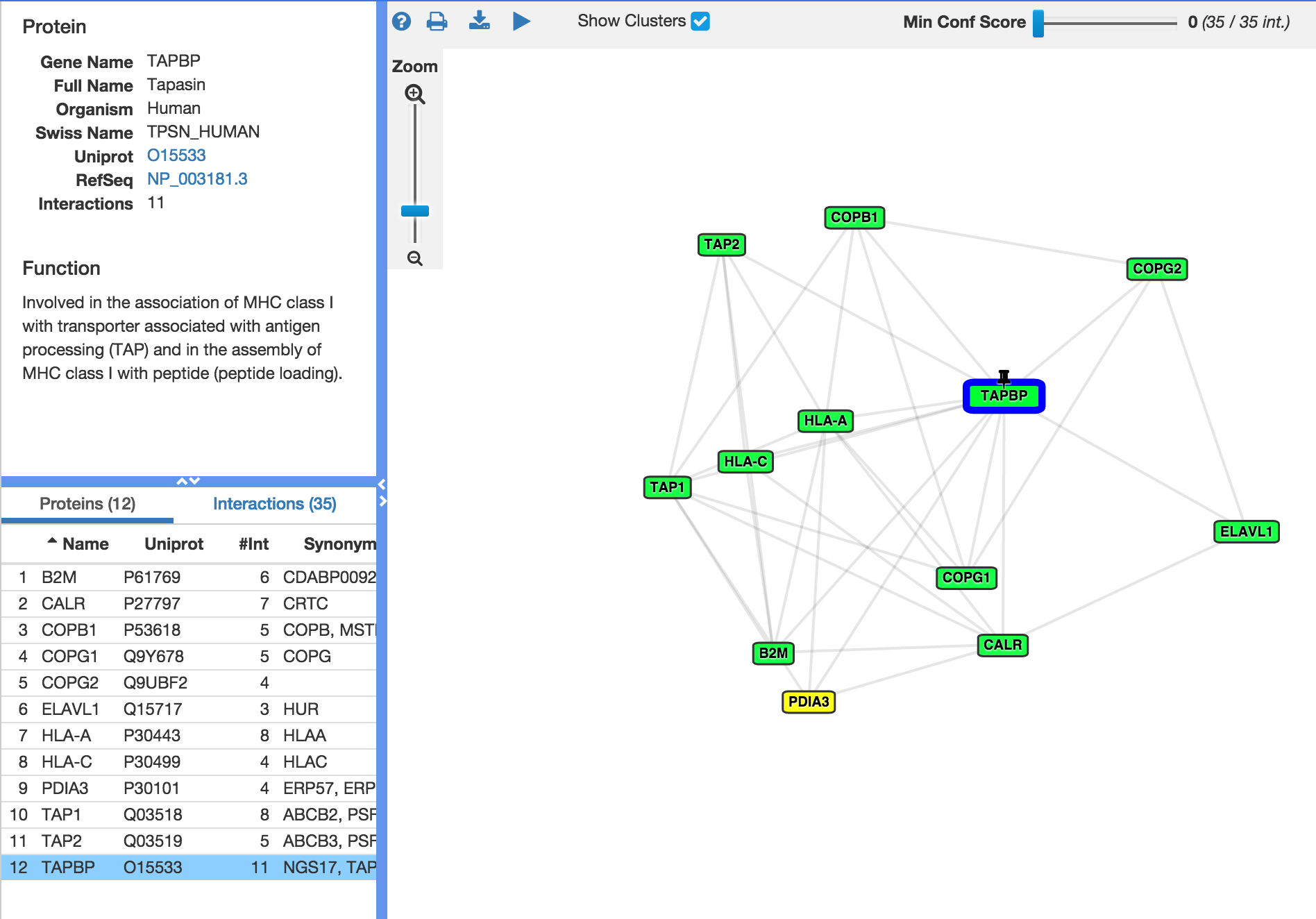
- VISUAL: Graphical protein browser - easy and intuitive.
- INTERACTIVE: The entire protein network is hyperlinked to information about proteins, functions, interactions and publications. Toggle between Protein or Interaction view by clicking nodes or edges in graph.
- EXPLORATORY: Expand any network by double clicking a protein in the visual display.
- EVIDENCE-BASED: Each interaction comes with a link to publications in chronological order on PubMed. Access the entire publication history for any interaction. Experimental techniques curated from the literature are listed for each interaction (e.g. Affinity Chromatography, X-ray crystallography).
- EXPANDABLE: Upload your own PPI or genetic/proteomic screen data and view in the context of published networks. Only you can see your data, by logging into your account.
- SHAREABLE: Link to your network with a simple URL (gpsprot.org/navigator.php?q=NAME, where NAME is gene name or UniProt ID) .
- EXPORTABLE: Export your network to a text file.
- PATHOGEN (HIV) PPIs included
Contact us at gpsprot@gmail.com.